Python中文网 - 问答频道, 解决您学习工作中的Python难题和Bug
Python常见问题
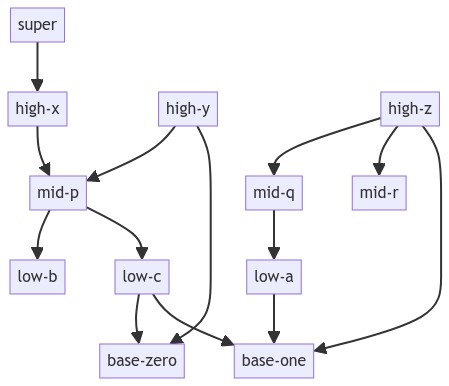
如果你看这个DAG(有向无环图):
我想创建一个dict,该dict将从最低节点到所有其他节点的距离映射到渲染图中每个节点距离底部的x位置(高度)。 对于给定的图形,它将是:
distance_nodes_map: {
0: {'base-zero', 'base-one'},
1: {'low-b', 'low-a', 'low-c'},
3: {'high-x', 'high-z', 'high-y'},
2: {'mid-r', 'mid-q', 'mid-p'},
4: {'super'}
}
我写了一个算法,对上面的图有效,但后来我测试了另一个图,它不再有效了。我尝试了一些算法和函数,比如最短路径或距离的子体,但我认为它们对计算距离没有帮助
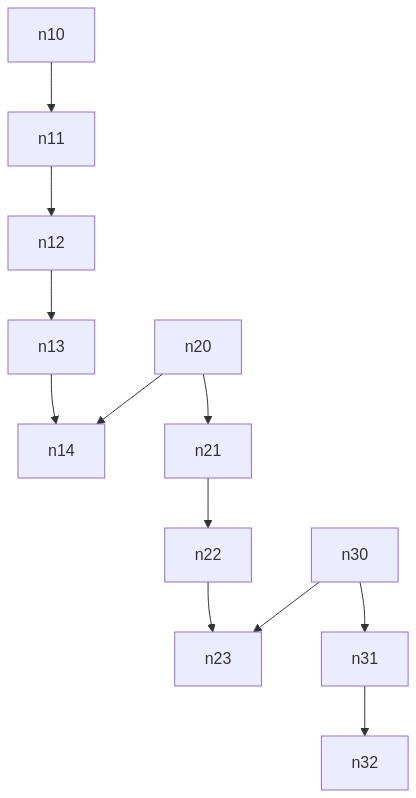
我的算法不适用于此图:
https://gist.github.com/timaschew/3b08a07243fa6f43773014ef5e705c96
以下是要点,包括:
- 一个python脚本,它读取YAML文件、依赖项/图形结构,并生成一个带有呈现的mermaid图形的HTML(我删除了以错误方式计算距离的算法)
- 此处显示的两个图形都是YAML文件
Tags: 文件算法图形距离yamlbase高度节点
热门问题
- 如何使用带Pycharm的萝卜进行自动完成
- 如何使用带python selenium的电报机器人发送消息
- 如何使用带Python UnitTest decorator的mock_open?
- 如何使用带pythonflask的swagger yaml将apikey添加到API(创建自己的API)
- 如何使用带python的OpenCV访问USB摄像头?
- 如何使用带python的plotly express将多个图形添加到单个选项卡
- 如何使用带Python的selenium库在帧之间切换?
- 如何使用带Python的Socket在internet上发送PyAudio数据?
- 如何使用带pytorch的张力板?
- 如何使用带ROS的商用电子稳定控制系统驱动无刷电机?
- 如何使用带Sphinx的automodule删除静态类变量?
- 如何使用带tensorflow的相册获得正确的形状尺寸
- 如何使用带uuid Django的IN运算符?
- 如何使用带vue的fastapi上载文件?我得到了无法处理的错误422
- 如何使用带上传功能的短划线按钮
- 如何使用带两个参数的lambda来查找值最大的元素?
- 如何使用带代理的urllib2发送HTTP请求
- 如何使用带位置参数的函数删除字符串上的字母?
- 如何使用带元组的itertool将关节移动到不同的位置?
- 如何使用带关键字参数的replace()方法替换空字符串
热门文章
- Python覆盖写入文件
- 怎样创建一个 Python 列表?
- Python3 List append()方法使用
- 派森语言
- Python List pop()方法
- Python Django Web典型模块开发实战
- Python input() 函数
- Python3 列表(list) clear()方法
- Python游戏编程入门
- 如何创建一个空的set?
- python如何定义(创建)一个字符串
- Python标准库 [The Python Standard Library by Ex
- Python网络数据爬取及分析从入门到精通(分析篇)
- Python3 for 循环语句
- Python List insert() 方法
- Python 字典(Dictionary) update()方法
- Python编程无师自通 专业程序员的养成
- Python3 List count()方法
- Python 网络爬虫实战 [Web Crawler With Python]
- Python Cookbook(第2版)中文版
您正在寻找一种绘制分层图的算法。有许多不同的算法,您应该选择最适合您需要的算法(例如,看看下面的论文A Technique for Drawing Directed Graphs,作者是Gansner等人。)
其中许多算法已经在Graphviz(一个非常著名和强大的图形可视化软件)中实现。一旦安装了它,就可以非常简单地计算您要查找的结果(
G是使用networkx.DiGraph构建的有向无环图):下面是几个使用您提供的数据的示例:
未经测试的伪代码,因为我的午休时间快结束了:
您有一个多根树,其中有一个选定的主根
对于每个根,创建一个子图,其中包含该根的所有可访问节点
从主根(根a)开始,计算对应子图a中所有节点到根的距离/最短路径长度
查找与主子图至少共享一个节点的所有子图,并选择与主根距离最小的节点(节点x)所在的子图(子图B)
计算子图b中所有节点到根b的距离。添加距离d(节点x,根a)。减去距离d(节点x,根b)
创建子图A和B的并集。重复步骤3-5,直到没有根为止
减去最大距离&;反转符号,使主根具有最大的距离/顺序值
编辑:
我的伪代码有效(*)。我责备用户的错误。;-)
收益率:
(*)“减去距离d(节点x,根b)”自然意味着节点x和根b之间的最长路径长度。很明显
相关问题 更多 >
编程相关推荐